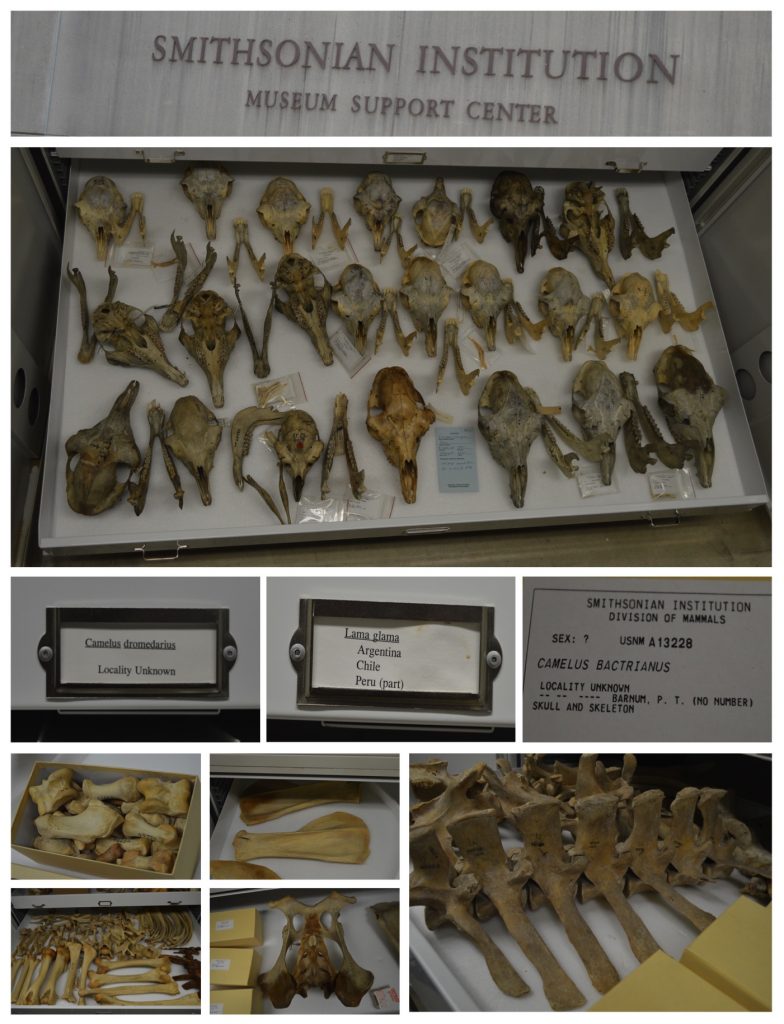
منذ أيام قليلة وعند بوابة المبنى الذي ألقي فيه محاضراتي رأيت إحدى طالباتي السابقات (نوال الشمري) مقبلة نحوي. كنت أظن أنها ترغب في إلقاء التحية كما هي عادة أغلب الطلبة الذين يعرفوني وأعرفهم. غير أن نوال فاجأتني بملف يحمل أوراق وقالت أنها أتت خصيصاً لهذا المبنى كي تعطيني هذا الملف. كان الملف يحوي ثلاث أعمال فنية تحاكي ما تعلمته نوال في مقرر المناعة والذي كان في الصيف الماضي. لم أتفاجأ بإبداع نوال خصوصاً وأنها في مقررين سابقين قدمت لي أعمال إبداعية مميزة. كانت المفاجأة أن العمل الإبداعي هو لمقرر لا أدرسه وفي غير تخصصي الدقيق.أسعدتني مبادرتها وأغرقني سعادةً أنها بعد اليوم لا تحتاجني كي تستمتع بما تتعلم في شتى المجالات. كانت تعتذر وتتساءل إن كنت أسمح باستقبال وتقييم أعمال من هذا النوع في خارج إطار ما أطرحه من مقررات وموضوعات. قلت لها وأقول لكل فرد مبدع يقرأ هذه الكلمات أنه شرف لي أن تشاركوني عقلكم وإبداعكم ويسعدني أن أضع أي عمل إبداعي في هذه الصفحة الإلكترونية و التي خصصتها لأعمال الطلبة المبدعين. لا شرط عندي ولا حد للإبداع أي كانت صوره. كانت على شكل رسومات أو كلمات وبأي لغة كانت. أي ابداع يكون خلاصة حب لعلم تعلمه الطلبة هو مرحب به وبشده. شكراً نوال